The Meijer Lab
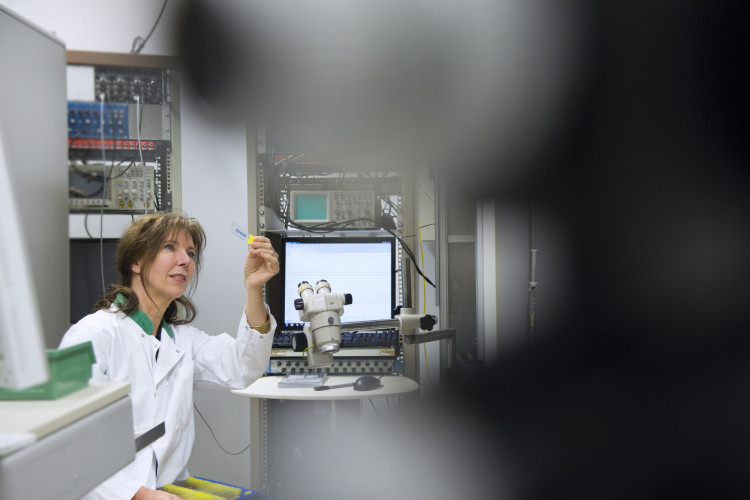

Circadian rhythms have evolved as essential adaptations to the temporal structure of our environment, governed by cycles of day and night and by seasonal variations in light. Prof. Johanna H. Meijer investigates how the brain integrates these external cues with intrinsic cellular signals, revealing how the fourth dimension—time—is encoded within neural circuits. Her work explores how the central nervous system continuously merges environmental factors and internal biological information into a coherent temporal program.
At the heart of her research lies a unique and self-developed in vivo methodology, which allows for direct, long-term recordings of neuronal activity in the brain’s biological clock—the suprachiasmatic nucleus (SCN)—under naturalistic conditions. By combining in vivo brain measurements with behavioral readouts such as sleep and physical activity, she was the first to simultaneously quantify how internal clock mechanisms and external behaviors influence one another in real-time. This approach has offered a level of resolution and realism unmatched in the field.
The Michel Lab
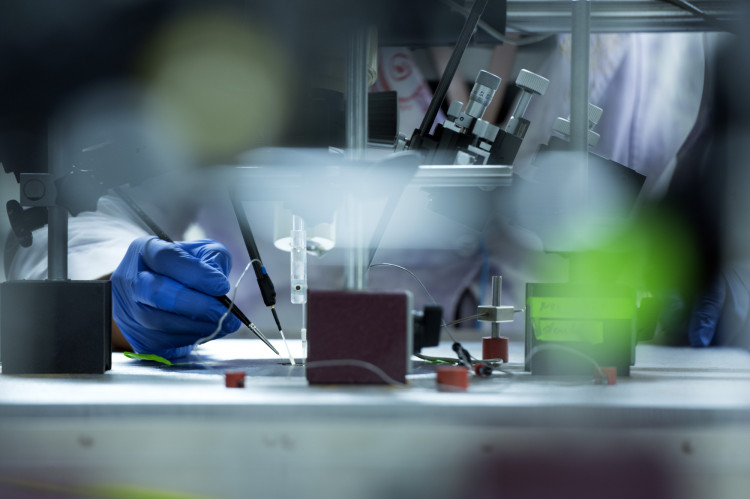

The study searches for potential interventions to maintain healthy brain function in the elderly. The aim of the project is to use a murine model to evaluate the impact of environmental light on aging brain circuits. Based on our recent findings that light regimes can influence the balance between excitatory and inhibitory neuronal activity, we will investigate how this affects neuronal network function. We want to use light as a tool to manipulate E/I balance in the old brain and evaluate the use light regimes to rectify ageing induced deficiency in brain function.
Among the many changes occurring in our body during aging, probably the least accessible to classical interventions are age-related alterations in brain function. Some of the reorganization taking place in the brain during aging may be as unavoidable and harmless as greying hair or may even be required to compensate for losses of neurons or synaptic contacts. Other changes however will lead to serious health problems like sleep disorders and may promote the development or progress of neurodegenerative diseases. These latter alterations in neuronal networks should be identified and be the focus of early interventions. In our own work, we demonstrated that the neuronal network of the Suprachiasmatic Nucleus (SCN), representing the central circadian clock in the hypothalamus, was altered in aging showing desynchronized neuronal activity which was correlated to fragmented sleep-wake patterns in old mice.
The De Boer Lab
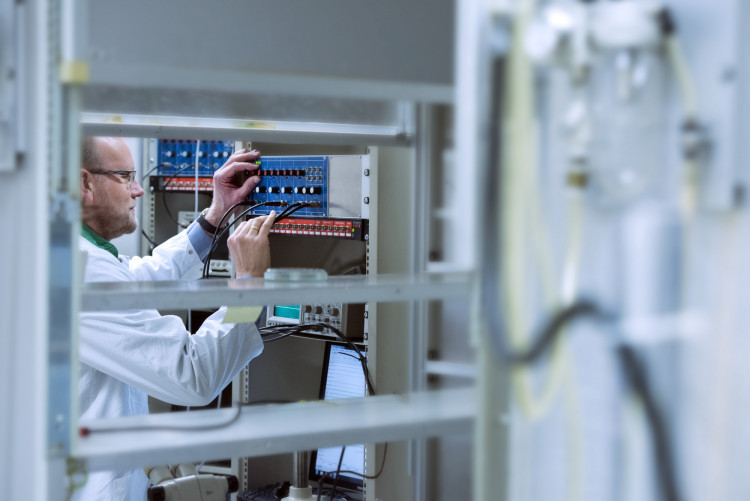

In the lab we combine different types of recording from rest-activity, to electroencephalogram, to electrophysiological recordings to investigate the interaction between the circadian clock, located in the suprachiasmatic nucleus of the hypothalamus, and sleep homeostatic processes.
Our research is inspired by the two process model of sleep regulation, the dominant model in sleep research since the beginning of the 1980’s. This model proposes that sleep pressure, or process S, increases during waking and decreases during sleep and that this pressure is reflected in the slow-waves (~2 Hz) of the electroencephalogram during non-rapid eye movement sleep. Next to this sleep homeostat there is the circadian clock which determines the timing of sleep. By recording these different variables, all related to sleep homeostasis and the circadian clock, and applying sleep deprivations protocols, light schedules or drugs that interfere with sleep or circadian clock functioning we try to elucidate the interaction between the two main processes that regulate sleep-wake behavior. Next to that we can use our techniques to investigate sleep and sleep regulation in different mouse models of human disease or apply newly developed pharmacological treatments to investigate their effect on sleep-wake behavior and brain functioning.
The Rohling Lab
.jpg)
The interaction between organs, brain regions, cells, but also between proteins and genes are pivotal for proper physiological functioning. These interactions can be described as a network of interacting agents. In network theory, a lot is known about the organization of networks and how this influences the outcomes of the interactions and the emergence of new functionality.
Brain networks are a hot topic as witnessed by the Human Brain Project of the EU, however, most brain network research is done at the macroscopic level using brain regions as the smallest entities, for example using fMRI, EEG or MEG data. Microscopic brain networks have the advantage of straightforward identification of the network nodes, being the cells. However, due to difficulty in data acquisition, these microscopic studies are seldom performed.
Our group investigates the emergent functional properties in the brain stemming from interaction in dynamical brain networks, and specifically how the microscopic networks in the brain shape brain-function. Currently, we explore multiple mathematical techniques to establish the functional connections between single neurons in the circadian clock, which is located in the suprachiasmatic nucleus (SCN), in order to create a network and apply network measures. Some of the techniques that we use are Detrended Fluctuation Analysis (DFA), clustering analysis, correlation analysis and oscillator analysis. The ultimate goal would be to understand the function of these dynamic networks in the brain, and possibly discover therapeutic treatments based on these network properties.